Rapid Activation of Catch-Bonds

Link to publication: https://doi.org/10.1021/acs.jctc.5c01181
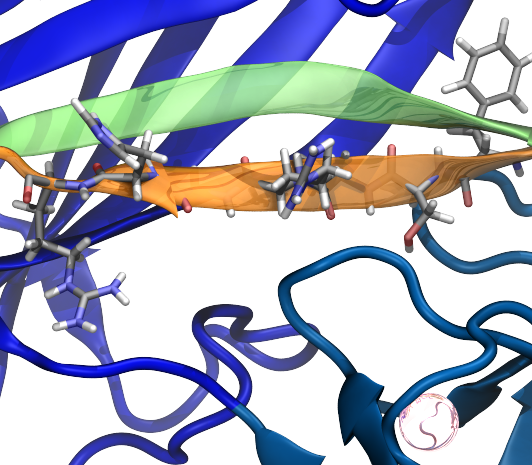
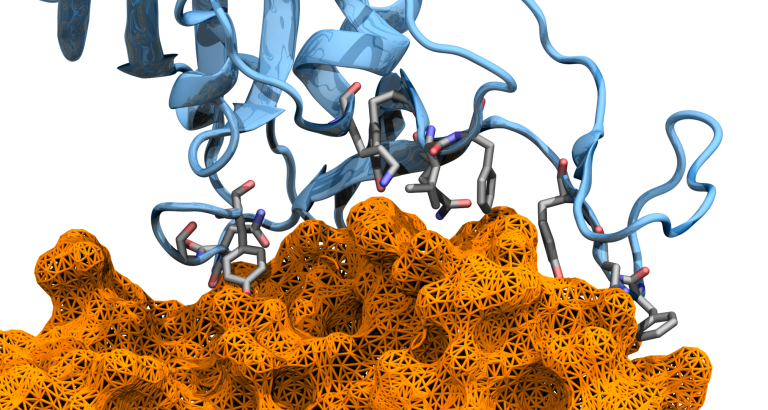
Mechanically resilient protein interactions are crucial for biological processes ranging from bacterial adhesion to human tissue formation. Catch-bonds, a unique class of protein interactions that strengthen under force, act like a molecular finger trap, tightening to prevent bond rupture. However, it remains unclear whether catch-bonds form immediately upon force application or require a specific force threshold for stabilization. Here, we employ an in silico single-molecule force spectroscopy approach that combines molecular dynamics (MD) simulations, dynamical network analysis, and AI-based modeling to investigate the XDoc:CohE complex, a hyperstable catch-bond found in cellulose-degrading bacteria. By analyzing amino acid interactions between XDoc and cohesin E, and between XDoc submodules (X-module and Doc), we show that AI regression models can accurately predict rupture forces using only short MD simulations, capturing key mechanostability features despite the binding interface’s complexity. Our results reveal that mechanostability signatures emerge early under force load, indicating that catch-bonds activate almost immediately. These findings provide new insights into the molecular principles governing force-dependent protein interactions and highlight the potential of AI-driven approaches for predicting and characterizing mechanostability, with broad implications for bioengineering and drug design.